BEST is a free phylogenetics program written by Liang Liu to estimate the joint posterior distribution of gene trees and species tree using multilocus molecular data that accounts for deep coalescence but not for other issues such as horizontal transfer or gene duplication. The program works within the popular Bayesian phylogenetics package MrBayes (Ronquist and Huelsenbeck, Bioinformatics, 2003). BEST parameters are defined using the prset command in MrBayes.
Development of the BEST project is a collaboration of Liang Liu, Dennis Pearl, and Scott Edwards.
The model:
BEST finds the joint posterior distribution of gene trees and the species tree for multi-locus data under a hierarchical Bayesian model. The BEST model assumes:
1. The prior distribution of species trees is uniform.
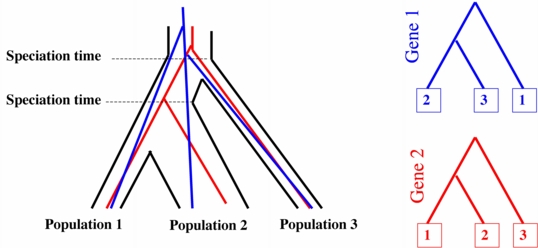
2. Given the species tree, the gene trees are conditionally independent and follow the coalescent model described by Rannala and Yang (Genetics, 2003).
3. Given the gene tree, the DNA sequences are conditionally independent of the species tree (for example, following one of the Markov substitution models available in MrBayes).
The algorithm:
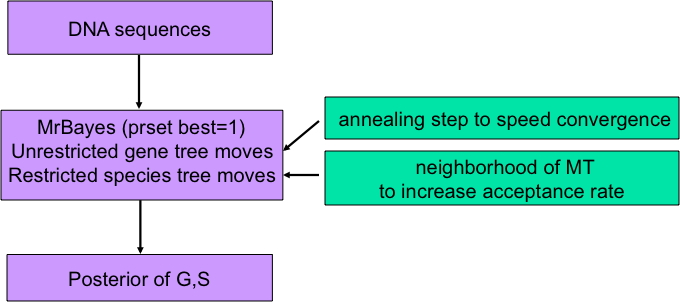
Proposal gene trees are made following the characteristics of the gene tree MCMC procedure specified by the user in MrBayes. This vector of gene trees is then paired with a species tree to form a joint (gene trees, species tree) proposal. The species tree is chosen from the space of species trees that fulfill the constraint that all divergences of species pairs must occur after the respective gene divergences occur. For a given set of gene trees this involves finding the Maximum Tree and modifying it at a random (Poisson) number of nodes while maintaining the defining constraint and making all compatible species trees attainable. Hastings ratios to define the acceptance probabilities are then calculated using the formulas found in Rannala and Yang (Genetics, 2003). Early in the MCMC, BEST uses an annealing step to down-weight the contribution of the prior in the Hastings ratio and move more quickly toward areas of high likelihood.